- Write structural formulas for all of the constitutionally isomeric compounds having the given molecular formula.
- C6H14
- C5H11Br
- C3H7N (13 isomers)
- Use the WebMO application on the C4 web server to complete the WebMO Tutorial (WebMO_Tutorial.PDF). Your login is "first-initial-last-name", all lower case with no spaces. Your password is "chem12".
As we learned in class, hybridization affects carbon-carbon bond lengths. In next few exercises we’ll use the WebMO application on the C4 web server to explore the carbon-carbon bond lengths of molecules containing double and triple bonds.
- Login to WebMO and build a model of 1,3-hexadiene (shown below).
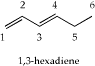
Be sure and use "comprehensive" or "comprehensive mechanics" clean-up to adjust the bond lenghts to within appropriate distances prior to conducting a geometry optimization. Once the molecule is constructed, conduct a geometry optimization using the AM1 theory level. View the completed job and inspect the carbon-carbon single bond lengths (C2 to C3, C4 to C5, and C5 to C6). Be sure to draw the molecule and indicate the bond lengths as part of your work to turn in.
- For comparison we are going to look at 1,3-hexadiyne (shown below).

Unfortunately this model is very difficult for the computer to handle. Becuase of the linear nature of the molecule, the geometry is difficult to define without knowing a lot about how the application works internally. I’m not exactly sure this is going to work for everyone, but we are going to try. I've built the model, optimized the geometry, and exported the job as a WebMO archive. You need to download the archive to your computer by clicking here (alkyne.tar). Save the archive titled "alkyne.tar" to your desktop or some place where you can easily find it.
From within WebMO Job Manager, click on the "New Job" button and hold. A pull down menu appears with the option of "New Job" or "Import." You want to select "Import."
In the first box type the name of the job. I recommend "1,3-hexadiyne." Then click on option 3 "Local WebMO archive - choose file." A window will pop-up allowing you to navigate to the location of the archive file (alkyne.tar) on your hard drive. Select the file and hit "Import Job." The window should reflect that the job was imported successfully. You can then hit "Return to Job Manager" to view the newly import job. The job should be at the top of the list in the Job Manager window.
View the completed job and inspect the carbon-carbon single bond lengths (C2 to C3, C4 to C5, and C5 to C6). Be sure to draw the molecule and indicate the bond lengths as part of your work to turn in.
- Look back at the values of the carbon-carbon bonds that you reported in problems 3 and 4. Explain the observed C-C bond distances in terms of the hybridization of each of the carbons in the bond. Your explanation should include reference to the sizes of the hybrid orbitals contributing to the bonds.
The electronegativity of carbon is 2.55. Numerous observations, however, indicate that sp-, sp2-, and sp3- hybridized carbon have slightly different electronegativities.
- Build models of ethane and ethylene (shown below).
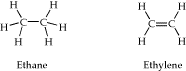
Be sure and use "comprehensive" or "comprehensive mechanics" clean-up to adjust the bond lengths to within appropriate distances prior to conducting a geometry optimization. Once the molecule is constructed, conduct a geometry optimization using the AM1 theory level. View the completed jobs. View the partial charges on the molecules. In which hydrocarbon is carbon most electronegative as measured by the extent to which it draws electrons away from an attached hydrogen? In which hydrocarbon is carbon the least electronegative?
To investigate acetylene you will have to import the completed model as we did in question 4. Download the archive of the model here (acetylene.tar). Once you have the model archive saved on your computer you can then import the model as before. Name the model acetylene. Compare the imported model of acetylene to the ethane and ethylene you built and optimized on your own. In answering the questions above, correlate hybridization with electronegativity of the carbons.
- Last one. Here we are going to look at the difference between Valence Bonding Theory and Molecular Orbital Theory. I want you to compare the two models below, ethylamine and acetamide.
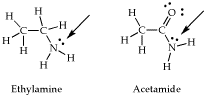
First I want you to make a prediction about the hybridization and geometry of the two nitrogen functional groups indicated in the picture.
Now I want you too look at the models of these two compounds. These are a little hard to model in WebMO, so I've included an archive that you can import here (nitrogen.tar), as we did before. This archive includes the two models so you only have to do the import once. These models also include the molecular orbitals for you to compare.
First, simply compare the geometries of the two nitrogen functional groups. Were they what you predicted? Next I want you too look at the Highest Occupied Molecular Orbital (HOMO) for each molecule. The loan pairs of electrons on a molecule generally occupy the HOMO. For the first molecule, ethylamine, the HOMO is molecular orbital 13. For the second molecule, acetamide, the HOMO is molecular orbital 16. Focus on the component of the orbital over the nitrogen atom in each molecule. Going back to Valence Bonding Theory, what type of orbital, sp-, sp2-, or sp3-, does the lone pair of electrons look like it resides in? Would Valence Bonding Theory have predicted this?